BrainVoyager 21 - BIDS, Defacing, 3D Viewer and Dark Mode
September 02, 2018
While version 21 of BrainVoyager has more visible changes of the user interface than many previous releases, the focus was on support for open standards, reproducible data pipelines and the foundation for using compute server backends. Furthermore, a content-focused dark interface and a new powerful graphics engine has been developed to further increase BrainVoyager's visualisation capabilities. The following sections shortly describe the main new features of the BrainVoyager 21.0 release that can be downloaded for 64-bit Windows, macOS and Linux operating systems. More details about the new features and improvements are provided in the release notes and the updated User’s Guide that can be downloaded as an e-book or viewed online in a web browser. Future updates of the 21 series will build on the foundation laid down in this first release to unlock remote computing capabilities and advanced visualisation features, such as integrated volume and surface rendering.
The created NIfTI file can then be directly opened with “File -> Open NIfTI” and used as the starting point for further processing. In addition to a document’s NIfTI file, an additional JSON file is stored with the same name (but with a “.json” file extension). The stored JSON file fulfils two purposes:
When specifying the location of the DICOM source data for your subjects, the data analysis manager in BIDS mode will automatically generate NIfTI files as well as JSON and TSV sidecar files instead of BV VMR and FMR-STC files. Furthermore, it will create a BIDS-compatible folder structure and places the created NIfTI files and sidecars in the respective folders using BIDS-compatible file names (see screenshot below for an example output of the data analysis manager). The folder structure of BIDS is rather similar to the old data analysis manger folder structure, placing subject folders containing session folders under a top-level project folder. The data within a session is, however further split according to their MRI scan type. For example, the "anat" folder contains anatomical data, while the "func" folder contains functional data of all runs of a session. The folder structure is partially also reflected in the BIDS file naming convention containing the subject name, session name, type of scan as well as the run number for functional data. For more details, download the complete BIDS specification.
BIDS is becoming a widely used standard in neuroimaging research collaboration and the dataset created by the data analysis manager is ready for sharing with colleagues as well as for uploading to (public) databases that support BIDS such as OpenNeuro. Note, however, that BIDS currently standardizes the folder structure and file naming for original scanner data but not for derived data, i.e., there is not yet a final recommendation how to organize preprocessed and (statistically) analyzed data on disk (a respective BIDS extension proposal is currently in development, see BIDS Extension Proposal 3). The current specification recommends to put derived data in the “derivatives” folder (see collapsed “derivatives” entry in the screenshot above) under the top-level project folder. When running preprocessing and analysis workflows in the updated data analysis manager, the generated output files will be stored in the derivatives folder creating for each workflow a separate sub-folder with a name reflecting the type of executed pipeline (e.g. functional preprocessing, spatial normalization). At present the output files are stored in conventional BrainVoyager formats but in future versions also the derived data will be saved as NIfTI plus sidecar files. In addition to the generated document files, each workflow folder contains a “parameters.json” file containing a human-readable description of parameters used by the respective workflow. The clear separation of workflows in separate folders and the human-readable description of parameters creates a transparent and reproducible system enabling easy processing of small and large datasets. The new data analysis manager also allows to create templates workflows for new projects ensuring consistent analysis across projects and lab members.
To deface original scanner data, BrainVoyager performs a temporary MNI normalization creating a 12-parameter affine spatial transformation matrix that is applied backwards to bring a defacing mask defined in MNI space into the data’s original (native) space where it is finally applied. This procedure ensures that no interpolation of the original voxel intensities is introduced, i.e., only the voxels inside the defacing mask are set to 0 while other voxels are untouched. In case of defacing DICOM files, the masked data is saved back — slice by slice — to the original DICOM files.
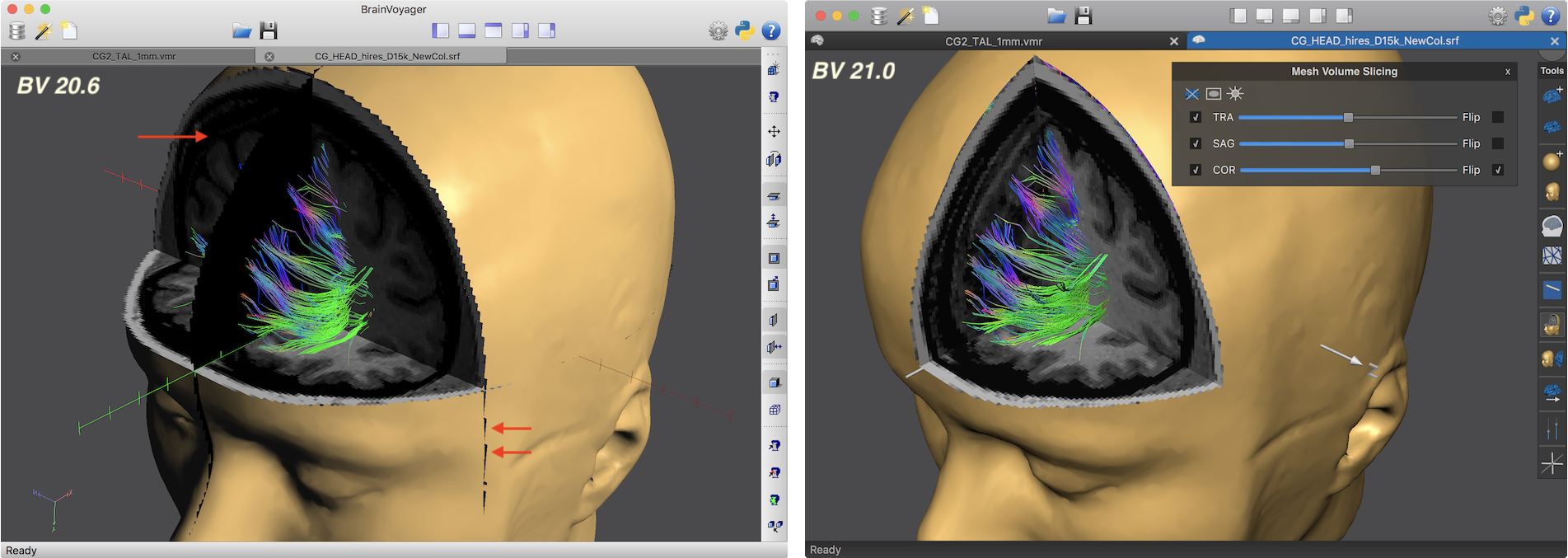
The screenshots above show another example highlighting the increased rendering quality in the new version (right side). The arrows on the left side point to artefacts produced in the old version along cuts and the inability to render correctly mesh cut-outs from three combined slice planes. Some new features of the 3D Viewer help to prepare figures for publication. When overlaying surface maps (SMPs), for example, the color bar of a selected map is shown in the right upper corner as in previous versions. In the new release (see screencast below), the panel is no longer fixed at this position but transformed into a movable panel when hovering the mouse pointer over the color bar. Clicking e.g. on the title bar of the panel allows moving the color bar to a more appropriate location in the 3D scene.
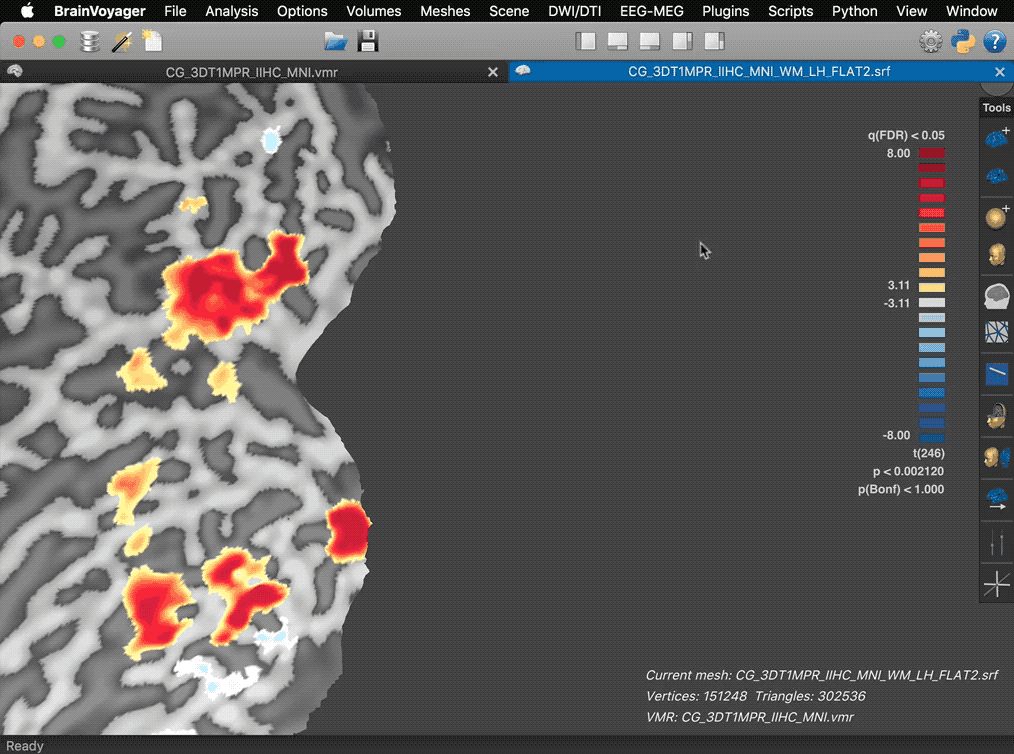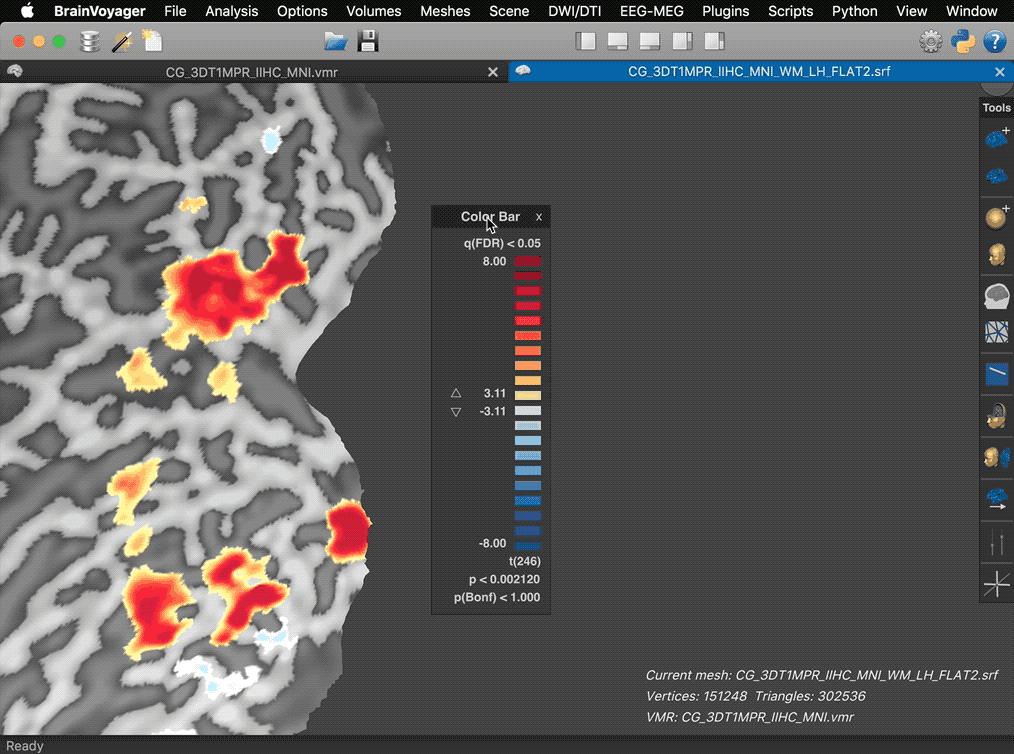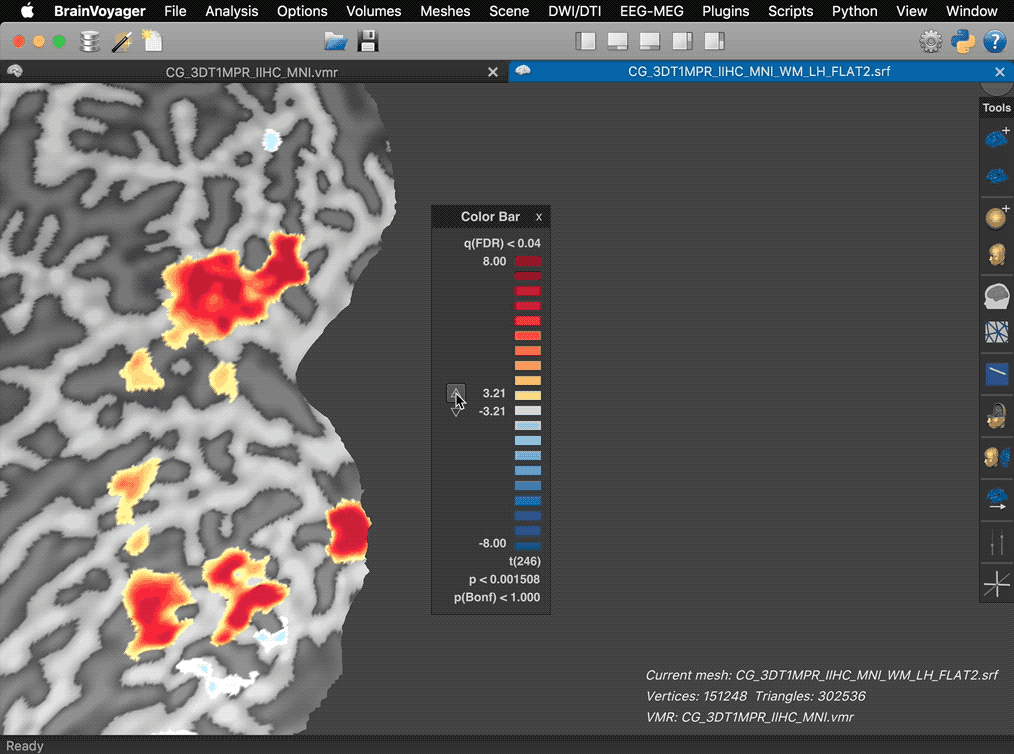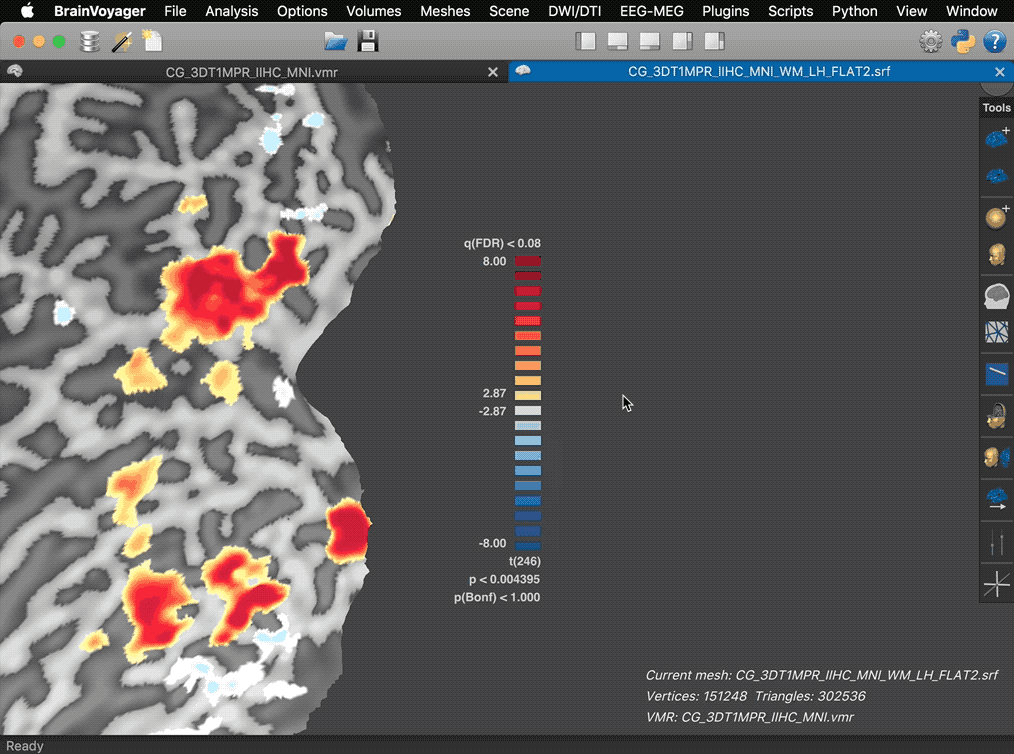
The panel also reveals action buttons for increasing and decreasing the current map threshold. When moving the mouse pointer out of the panel, the action buttons and panel frame disappear automatically leaving the color bar ready for saving screenshots for figures.
Besides the new features highlighted here, there are many other enhancements in this release, including an improved Python coding environment (now using Python 3.6 instead of 2.7 and first time support for Linux), MNI support for the Granger Causality and Psycho-Physiological Interaction (PPI) plugin that is now a core plugin installed directly with the software. We think that you will like the new release and we are looking forward to your feedback and suggestions for further improvements of our flagship software product!
DICOM-to-NIfTI Conversion and BIDS Compatibility
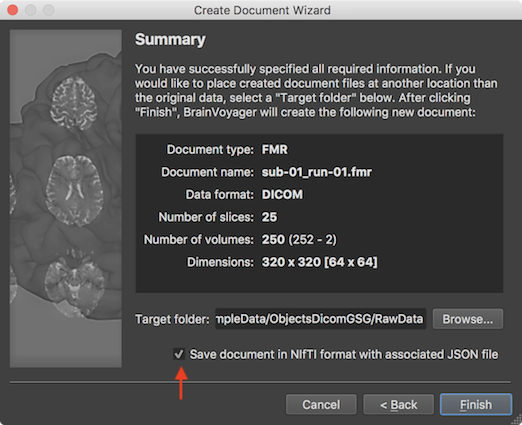
BrainVoyager 20 started to make NIfTI files first-class citizens. This new release continues this process by adding a “Save NIfTI” item directly in the “File” menu. Furthermore, the “Create Document Wizard” now provides the option to save created documents in the NIfTI format, i.e. it offers DICOM-to-NIfTI conversion (see arrow in screenshot below):
The created NIfTI file can then be directly opened with “File -> Open NIfTI” and used as the starting point for further processing. In addition to a document’s NIfTI file, an additional JSON file is stored with the same name (but with a “.json” file extension). The stored JSON file fulfils two purposes:
- it stores relevant information from the DICOM header in a human-readable form, which is also required for BIDS-compatibility (see below)
- it contains a “BrainVoyagerInfo” entry (JSON object) that stores information that is specific to BrainVoyager, i.e. information that is usually stored in the header of functional (FMR) and anatomical (VMR) document files and for which the NIfTI header alone is not sufficient.
BIDS Folder Structure and updated Data Analysis Manager
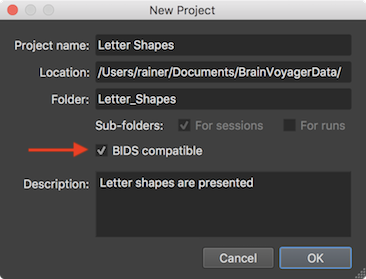
When creating a new project, the BrainVoyager data analysis manager will now operate “in BIDS mode” as default (see arrow in screenshot below). It is still possible to enable the old folder structure but its usage is discouraged.
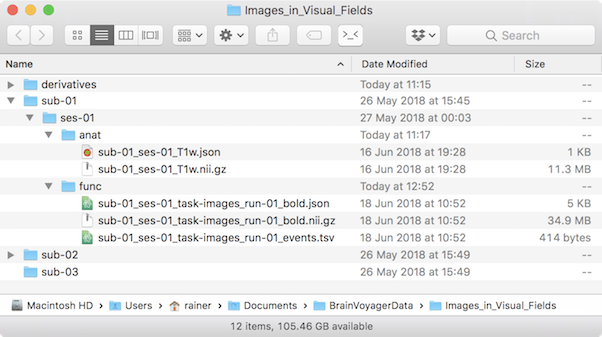
When specifying the location of the DICOM source data for your subjects, the data analysis manager in BIDS mode will automatically generate NIfTI files as well as JSON and TSV sidecar files instead of BV VMR and FMR-STC files. Furthermore, it will create a BIDS-compatible folder structure and places the created NIfTI files and sidecars in the respective folders using BIDS-compatible file names (see screenshot below for an example output of the data analysis manager). The folder structure of BIDS is rather similar to the old data analysis manger folder structure, placing subject folders containing session folders under a top-level project folder. The data within a session is, however further split according to their MRI scan type. For example, the "anat" folder contains anatomical data, while the "func" folder contains functional data of all runs of a session. The folder structure is partially also reflected in the BIDS file naming convention containing the subject name, session name, type of scan as well as the run number for functional data. For more details, download the complete BIDS specification.

BIDS is becoming a widely used standard in neuroimaging research collaboration and the dataset created by the data analysis manager is ready for sharing with colleagues as well as for uploading to (public) databases that support BIDS such as OpenNeuro. Note, however, that BIDS currently standardizes the folder structure and file naming for original scanner data but not for derived data, i.e., there is not yet a final recommendation how to organize preprocessed and (statistically) analyzed data on disk (a respective BIDS extension proposal is currently in development, see BIDS Extension Proposal 3). The current specification recommends to put derived data in the “derivatives” folder (see collapsed “derivatives” entry in the screenshot above) under the top-level project folder. When running preprocessing and analysis workflows in the updated data analysis manager, the generated output files will be stored in the derivatives folder creating for each workflow a separate sub-folder with a name reflecting the type of executed pipeline (e.g. functional preprocessing, spatial normalization). At present the output files are stored in conventional BrainVoyager formats but in future versions also the derived data will be saved as NIfTI plus sidecar files. In addition to the generated document files, each workflow folder contains a “parameters.json” file containing a human-readable description of parameters used by the respective workflow. The clear separation of workflows in separate folders and the human-readable description of parameters creates a transparent and reproducible system enabling easy processing of small and large datasets. The new data analysis manager also allows to create templates workflows for new projects ensuring consistent analysis across projects and lab members.
Defacing 3D Anatomical Data
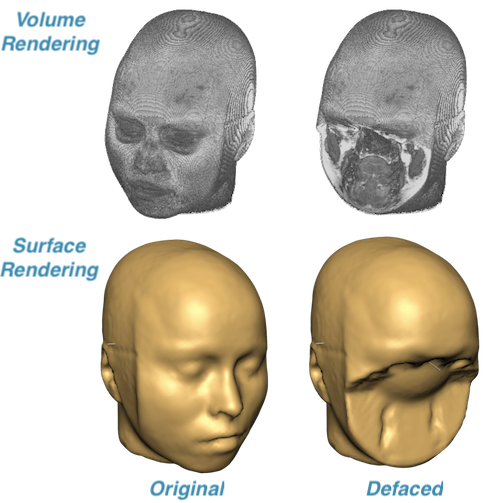
An advantage of using NIfTI files for raw data instead of DICOM files is that the small header of NIfTI files makes it easy to ensure that the data are anonymized. The BIDS folder can thus be shared without violating data privacy. There is one issue though, namely that the identity of participants may be revealed by reconstructing the skin surface from 3D anatomical data. To remedy this, BrainVoyager 21 comes with defacing tools (see screenshot below) that operate on created VMR documents (that can be saved as NIfTI) but defacing can also be applied to the respective DICOM files themselves; in combination with BrainVoyager’s DICOM header and file name anonymization tools, this is useful in case you need to share your scanner data in DICOM format.
To deface original scanner data, BrainVoyager performs a temporary MNI normalization creating a 12-parameter affine spatial transformation matrix that is applied backwards to bring a defacing mask defined in MNI space into the data’s original (native) space where it is finally applied. This procedure ensures that no interpolation of the original voxel intensities is introduced, i.e., only the voxels inside the defacing mask are set to 0 while other voxels are untouched. In case of defacing DICOM files, the masked data is saved back — slice by slice — to the original DICOM files.
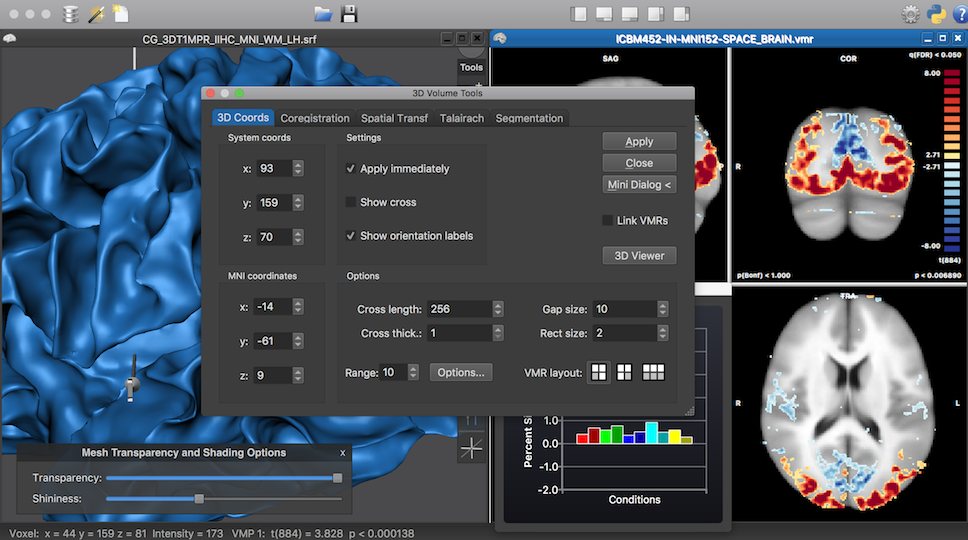
Content-Focused Dark Interface
BrainVoyager 21 comes with a unified cross-platform dark theme for its interface that is enabled as default. The dark interface (see screenshot below and dialogs at the start of this blog post) not only looks good but also supports a content-focused working style. If one wants not to use the dark interface, or only at certain times (e.g. evening), one can switch to the classic interface with native user elements and colors of the respective operating system. The embedded panels of the new 3D Viewer (see next section) support at present, however, only the dark interface.
State-Of-The-Art 3D Rendering Engine
The original 3D rendering engine of BrainVoyager was built in the end of the 1990’s using version 1.1. of OpenGL. While the 3D visualisation possibilities have been improved over the years, they were built on top of the — now old-style and inefficient — original OpenGL programming approach. In the meantime OpenGL has become much more powerful and flexible exploiting the power of modern 3D graphics cards (GPU’s) much more efficiently by using so-called shaders, which are small programs that are loaded and executed directly on the GPU. This programmable shader approach forms also the basis of currently developed successors of OpenGL like Vulkan and Metal (specific to macOS). The new graphics engine of BrainVoyager 21 has adopted the latest programmable graphics technology of OpenGL and beyond providing highly efficient 3D rendering that is substantially faster (up to 10x) than the older OpenGL approach, Furthermore, the new rendering engine provides sophisticated specialised rendering effects such as 3D texture sampling and integrated volume rendering that will be described in subsequent blog posts.
New 3D Viewer
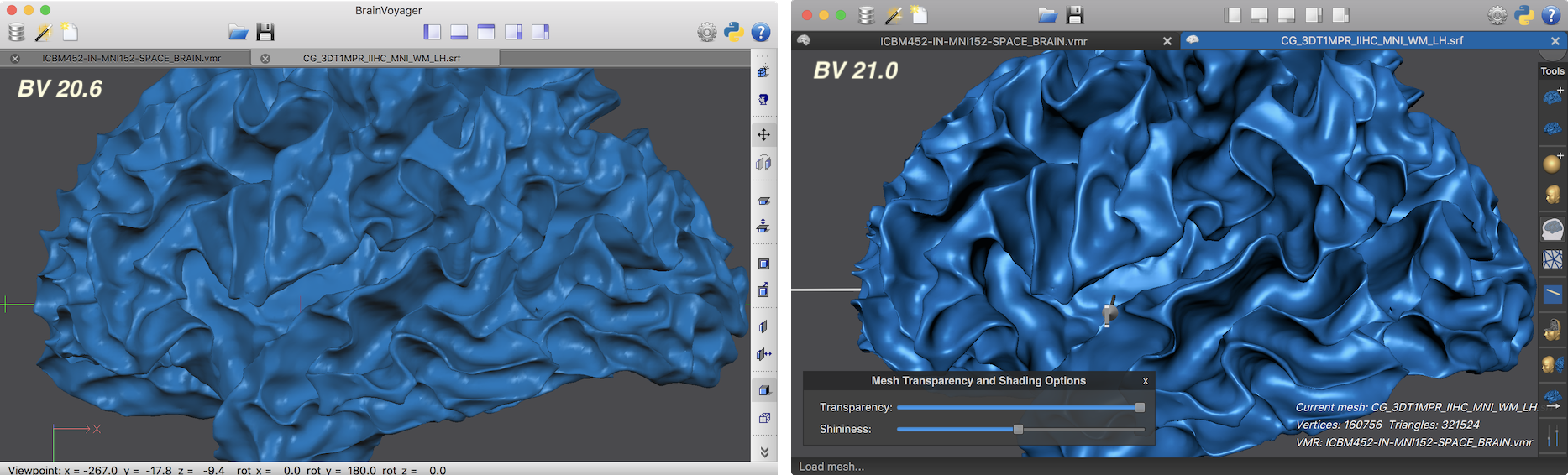
The new rendering engine is the core component of the new 3D Viewer of BrainVoyager 21 (see below) replacing the old “surface module window”. The 3D Viewer supports touch displays on Windows and track pads on macOS supporting the use of one-finger rotation and two-finger pinch-zoom and translation gestures. The 3D Viewer has a new toolbar that can be used, as in previous versions, to trigger important actions, such as mesh creation and surface reconstruction. It also invokes new floating panels that provide a more flexible and efficient user interface to handle scenes of multiple 3D models. The screenshot above shows a typical rendering gin the previous BrainVoyager version (20.6) and on the right side the same brain surface as it appears in the new 3D Viewer with the new toolbar and one floating panel for changing transparency and "shininess" of the mesh.
The screenshots above show another example highlighting the increased rendering quality in the new version (right side). The arrows on the left side point to artefacts produced in the old version along cuts and the inability to render correctly mesh cut-outs from three combined slice planes. Some new features of the 3D Viewer help to prepare figures for publication. When overlaying surface maps (SMPs), for example, the color bar of a selected map is shown in the right upper corner as in previous versions. In the new release (see screencast below), the panel is no longer fixed at this position but transformed into a movable panel when hovering the mouse pointer over the color bar. Clicking e.g. on the title bar of the panel allows moving the color bar to a more appropriate location in the 3D scene.

The panel also reveals action buttons for increasing and decreasing the current map threshold. When moving the mouse pointer out of the panel, the action buttons and panel frame disappear automatically leaving the color bar ready for saving screenshots for figures.
Besides the new features highlighted here, there are many other enhancements in this release, including an improved Python coding environment (now using Python 3.6 instead of 2.7 and first time support for Linux), MNI support for the Granger Causality and Psycho-Physiological Interaction (PPI) plugin that is now a core plugin installed directly with the software. We think that you will like the new release and we are looking forward to your feedback and suggestions for further improvements of our flagship software product!